Research
There is an abundance of publicly available data about various biological systems, but it can be difficult to draw insight from individual datasets. Our lab develops algorithms that integrate these data to help model and understand complex biological systems. Doing this allows us to investigate many different biological conditions, including those with limited data, such as rare diseases. We recognize that our lab won’t have all the answers, or even all of the questions, so we aim to develop tools and processes that any biologist can reuse. Our approach to research prioritizes transparency, rigor, and reproducibility.
The citations on this page were generated automatically from just identifiers using the Manubot cite utility developed right here in the Greene Lab!
2024

2023























2022






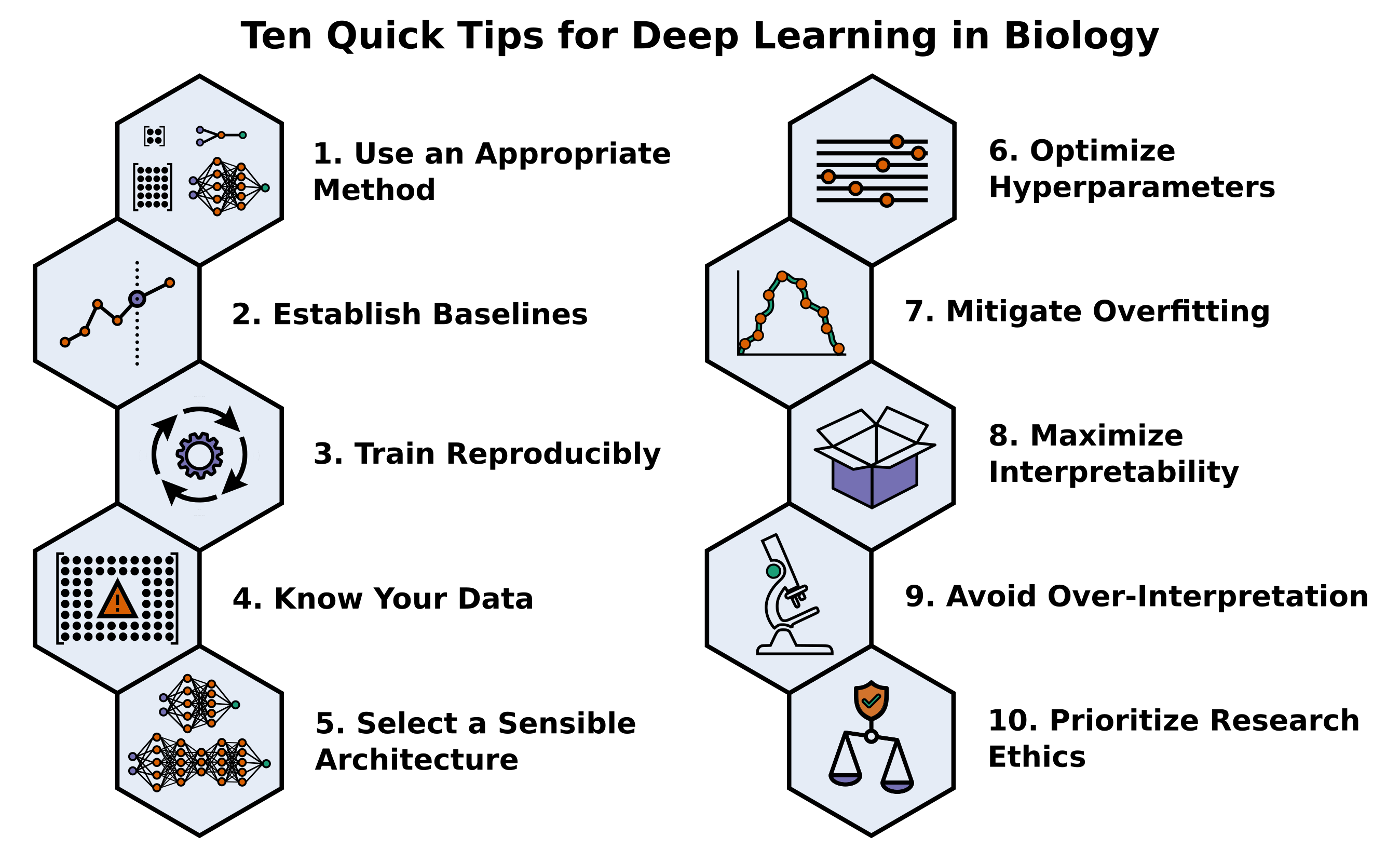




2021








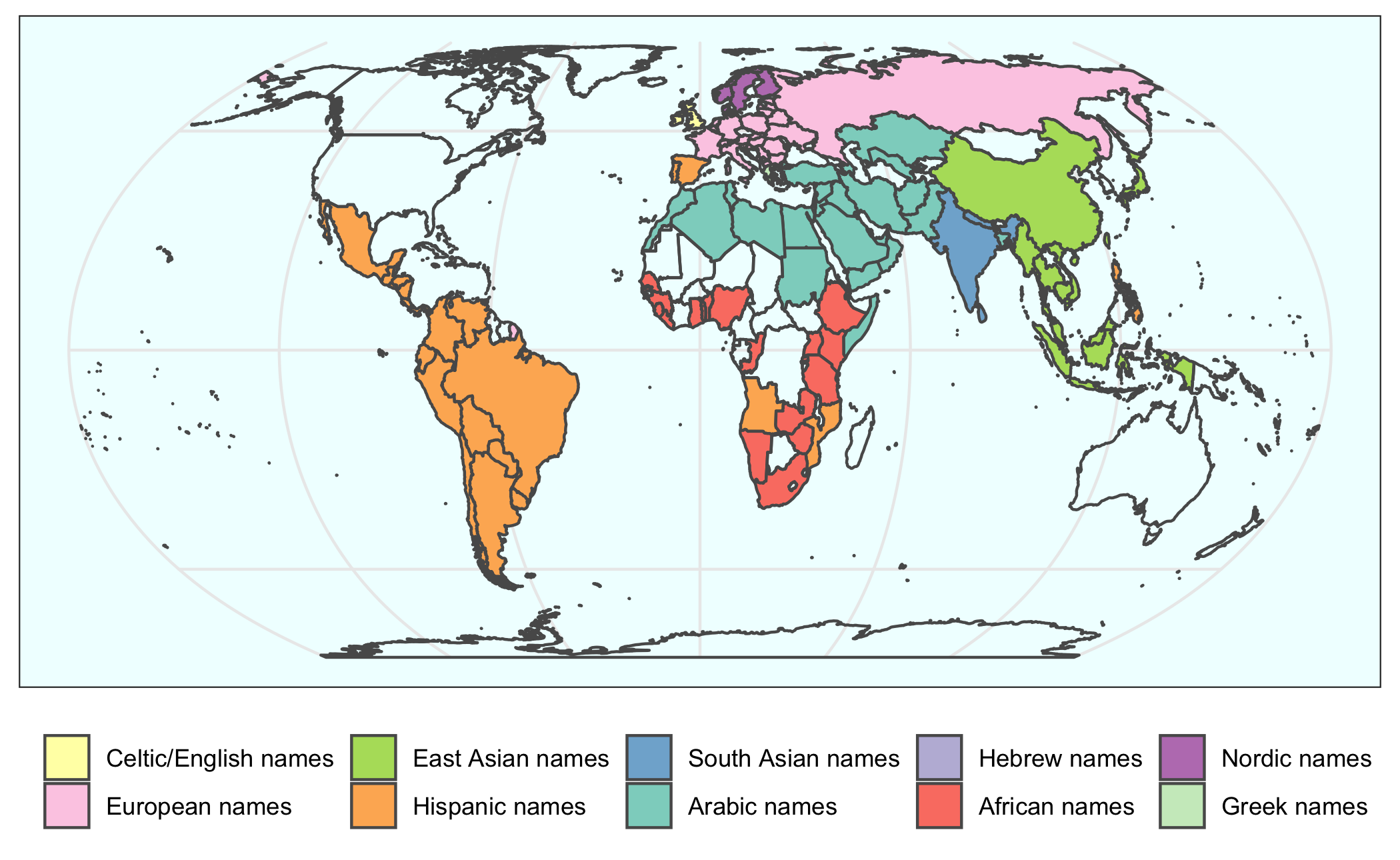


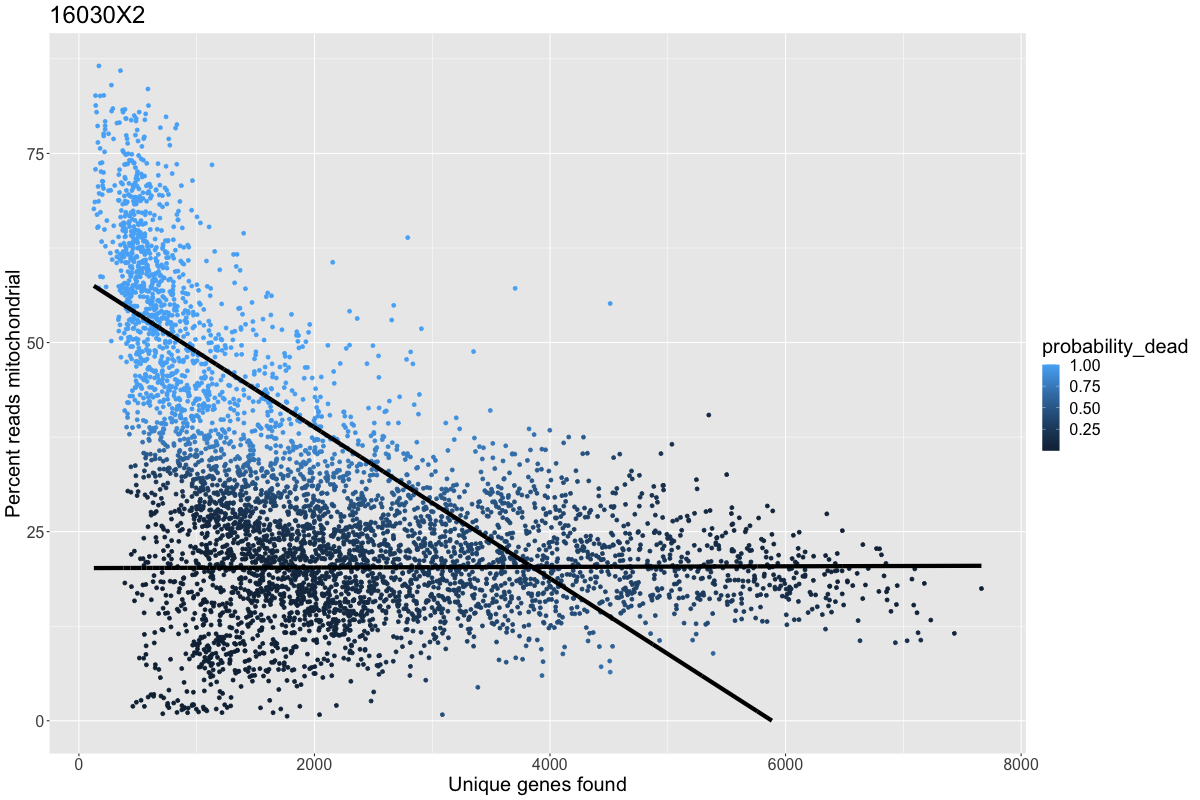
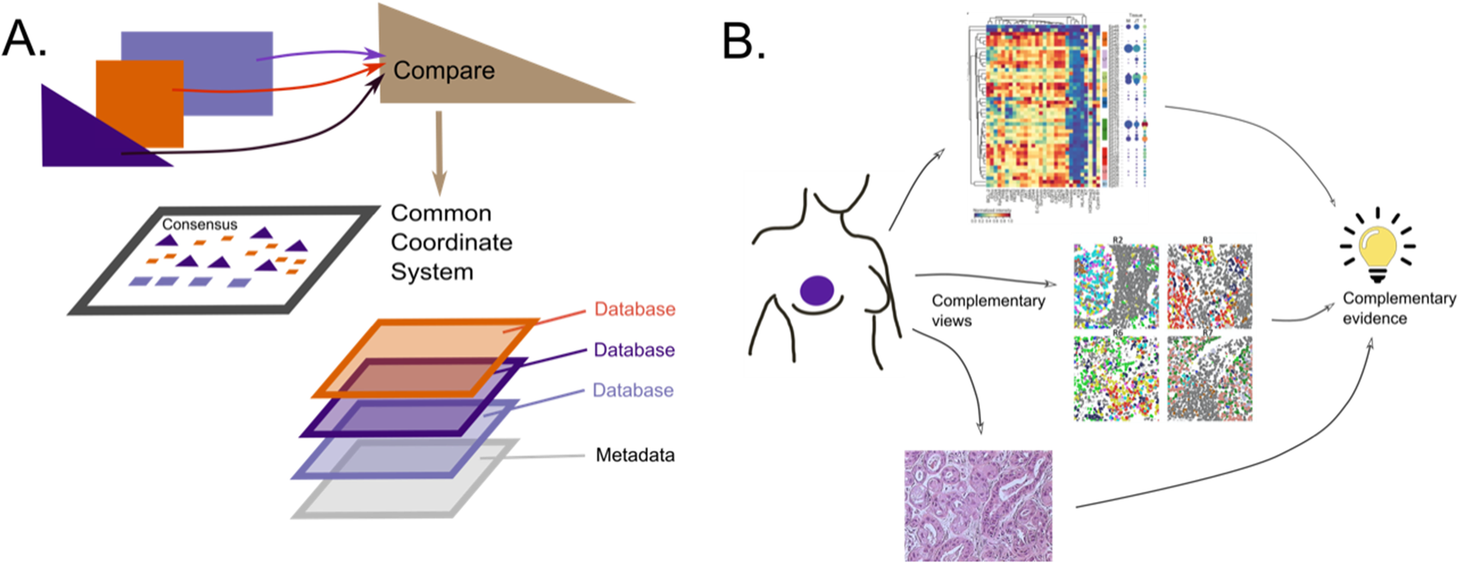








2020















2019












2018













2017








2016














2015





2014

2013


2012

